New insights into how the immune system recognizes viral RNA
Researchers clarify interplay between two proteins involved in viral detection, paving the way for improved mRNA vaccines and antiviral therapies
Study reveals how two proteins cooperate in a key early step of antiviral detection, as reported by researchers at Science Tokyo. Using cryo-electron microscopy and high-speed atomic force microscopy, they found that LGP2 binds to viral RNA and recruits MDA5 molecules, as if threading beads on a string. This creates a scaffold that facilitates the formation of a large signaling complex, which ultimately triggers an innate immune response.
How Sensor Proteins Cooperate to Detect Viral RNA Inside Cells
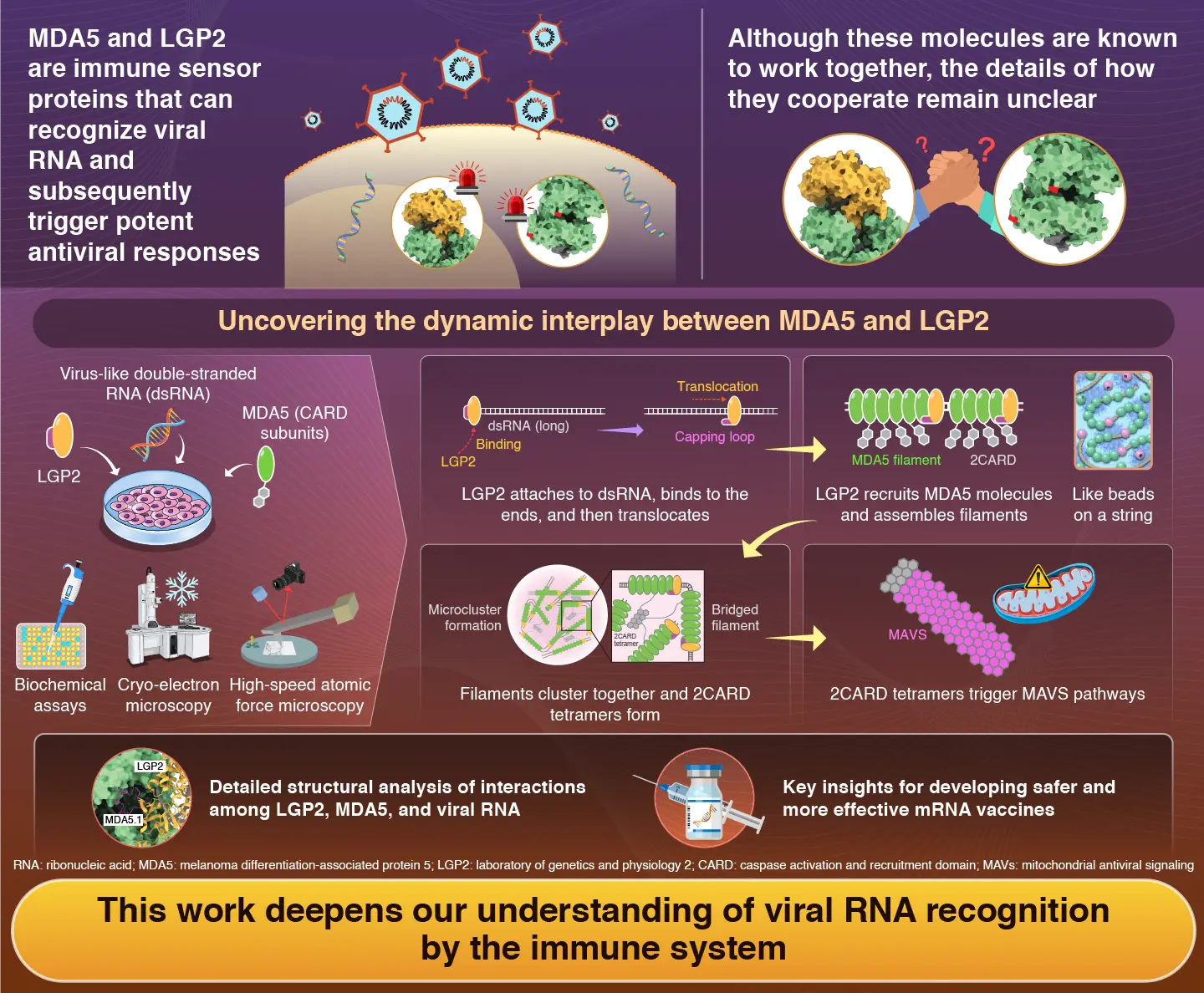
The innate immune system acts as the body's first line of defense against viral infections. When viruses enter cells, they often produce double-stranded ribonucleic acid (dsRNA) as part of their replication process. Specialized immune receptors present in the cell's cytoplasm are constantly searching for dsRNA as a sign of infection. One such receptor, melanoma differentiation-associated protein 5 (MDA5), plays a central role by recognizing viral dsRNA and forming filament-like structures along it. Once enough MDA5 molecules assemble, they trigger a signaling cascade that ultimately leads to the production of antiviral compounds.
Another important protein in this process is laboratory of genetics and physiology 2 (LGP2). LGP2 binds to viral RNA and moves along it through an energy-dependent process known as translocation. Although LGP2 cannot directly send antiviral signals on its own, scientists have long known that LGP2 and MDA5 cooperate to detect viral infections. However, exactly how LGP2 recognizes viral RNA and assists MDA5 has remained unclear.
To address this question, a research team led by Associate Professor Kazuki Kato (tenure-track) from the Mechanistic Immunology Research Unit, Institute of Integrated Research, Institute of Science Tokyo (Science Tokyo), Japan, together with Professor Osamu Nureki and graduate student Nina Kurihara (at the time of the study) from the Department of Biological Sciences, Graduate School of Science, The University of Tokyo, Japan, investigated the details of this molecular partnership. Using a combination of biochemical experiments and advanced imaging techniques, the team examined how LGP2 and MDA5 interact with viral RNA and with each other. Their findings were published online in the journal Molecular Cell on January 19, 2026.
The researchers first showed that LGP2 is especially important when viral RNA molecules are relatively short. On its own, MDA5 responds best to long dsRNA strands. However, when LGP2 was present, MDA5 could efficiently form short filaments even on shorter RNA molecules, activating downstream antiviral signaling.
To understand how this happens at the molecular level, the team used cryo-electron microscopy to probe the proteins' structure and high-speed atomic force microscopy to visualize them in action. They discovered that LGP2 first binds to the ends of a dsRNA molecule. Then, it uses energy from ATP to move along the RNA strand. As it advances, LGP2 functions as a scaffold, recruiting MDA5 molecules behind it and helping them assemble into stable filament structures. The researchers draw an analogy to threading beads on a string: the string represents dsRNA, while LGP2 acts as the leading bead of a group of MDA5 beads.
The team also found that LGP2 ultimately promotes the formation of small clusters of MDA5 filaments. These clusters enhance the activation of mitochondrial antiviral signaling (MAVS), a key signaling protein that amplifies antiviral responses inside cells. "Our study clarifies the precise mechanisms by which LGP2 recognizes viral RNA and cooperates with MDA5 to activate immune responses," says Kato.
By clarifying how LGP2 and MDA5 work together, this study provides a detailed picture of a key early step in antiviral defense. Such knowledge could guide future efforts to fine-tune immune responses in antiviral therapies and RNA-based medical technologies, as Kato concludes: "These findings deepen our understanding of viral RNA recognition and are expected to contribute to the design of safer and more effective mRNA vaccines."
Reference
- Authors:
- Nina Kurihara1,2, Yukari Isayama1, Jiayan Zhang1, Takashi Yamashita3, Kentaro Awaji3, Yukiko Ito3, Ayumi Yoshizaki3, Takahisa Kouwaki4, Hiroyuki Oshiumi4, Hiroshi Nishimasu5,6,7, Mikihiro Shibata8,9, Osamu Nureki2*, and Kazuki Kato1*
- Title:
- Molecular mechanism of MDA5 nucleation and filament formation by LGP2
- Journal:
- Molecular Cell
- Affiliations:
- 1Mechanistic Immunology Research Unit, Institute of Integrated Research, Institute of Science Tokyo, Japan
2Department of Biological Sciences, Graduate School of Science, The University of Tokyo, Japan
3Department of Dermatology, Graduate School of Medicine, The University of Tokyo, Japan
4Department of Immunology, Graduate School of Medical Sciences, Faculty of Life Sciences, Kumamoto University, Japan
5Structural Biology Division, Research Center for Advanced Science and Technology, The University of Tokyo, Japan
6Department of Chemistry and Biotechnology, Graduate School of Engineering, The University of Tokyo, Japan
7Inamori Research Institute for Science, Japan
8Institute for Frontier Science Initiative, Kanazawa University, Japan
9WPI Nano Life Science Institute (WPI-NanoLSI), Kanazawa University, Japan
Related articles
Further information
Tenure-track Associate Professor Kazuki Kato
Mechanistic Immunology Research Unit, Institute of Integrated Research, Institute of Science Tokyo
Contact
Public Relations Division, Institute of Science Tokyo
- media@adm.isct.ac.jp
- Tel
- +81-3-5734-2975